
 Back to homepage
Back to homepage
The Laboratory
Capabilities based on dedicated technical platforms and high-quality partners
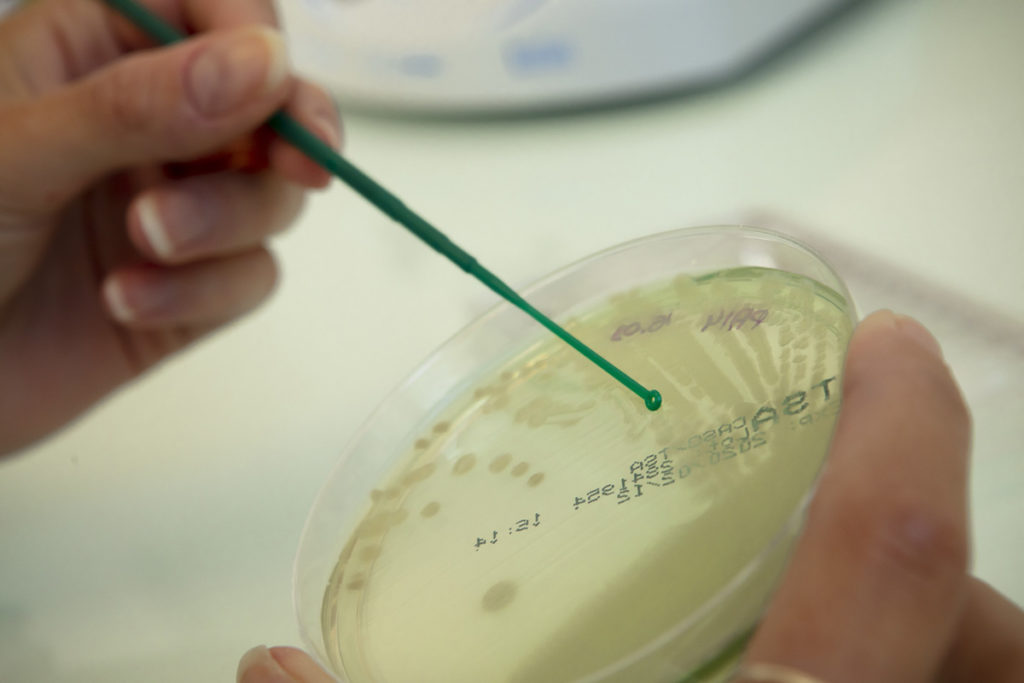
MICROBIOLOGY
Our microbiology platform allows us to handle microorganisms of various types: bacteria, yeast, molds, and phages, including a large number of pathogenic strains. Besides, the acquisition of an anaerobic station is a crucial element, allowing us to cultivate strains that have demanding atmospheric conditions.
In summary, the microbiology platform includes:
- 125 m² laboratory complying with Biosafety Level 2 (BSL2)
- Anaerobic station
- Analytical systems such as Tecan Spark 10M or BMG Labtech Spectrostar Nano microplate readers
- Microorganism counting systems (Bactobox and Spiral platter Eddy Jet)
- Electroporators
- Ultracentrifuge
- Climate-controlled chambers
- Remote Bruker® Maldi-Tof mass spectrometry platform
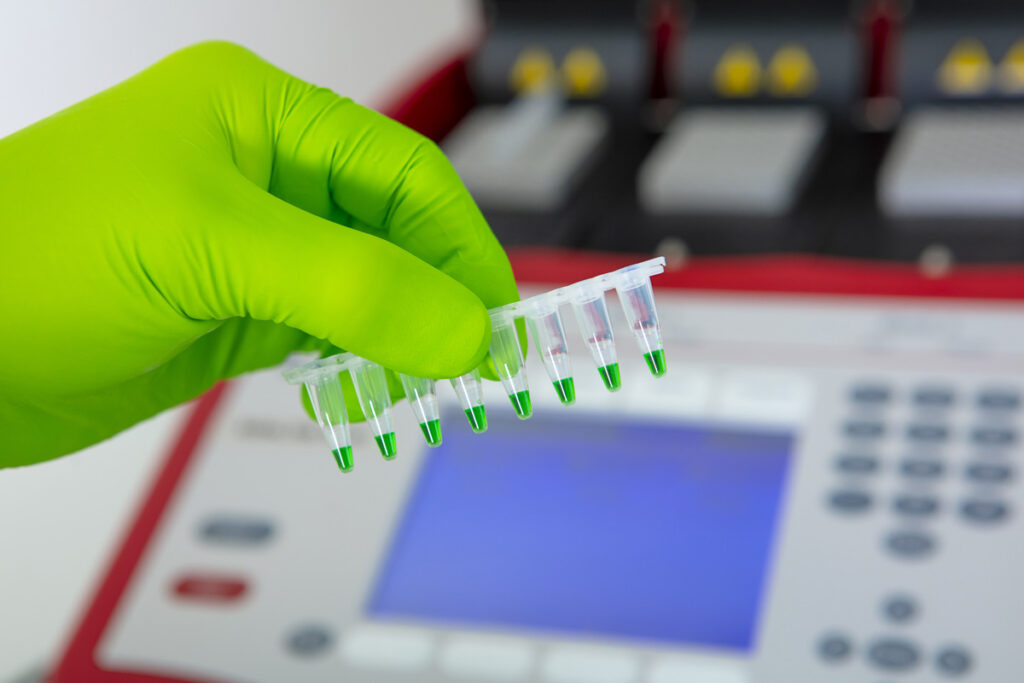
MOLECULAR BIOLOGY
Our molecular biology platform allows us to perform various missions such as genetic engineering or gene expression studies. This molecular biology technical platform complements the microbiology platform, as projects often require the use of both capabilities.
In summary, the molecular biology platform includes:
- 46 m² laboratory complying with Biosafety Level 2 (BSL2)
- Biometra Trio 48 Thermocyclers
- Real-time PCR systems
- Electrophoresis and DNA visualization systems
- Nucleic acid quantification systems
- Climate-controlled chambers
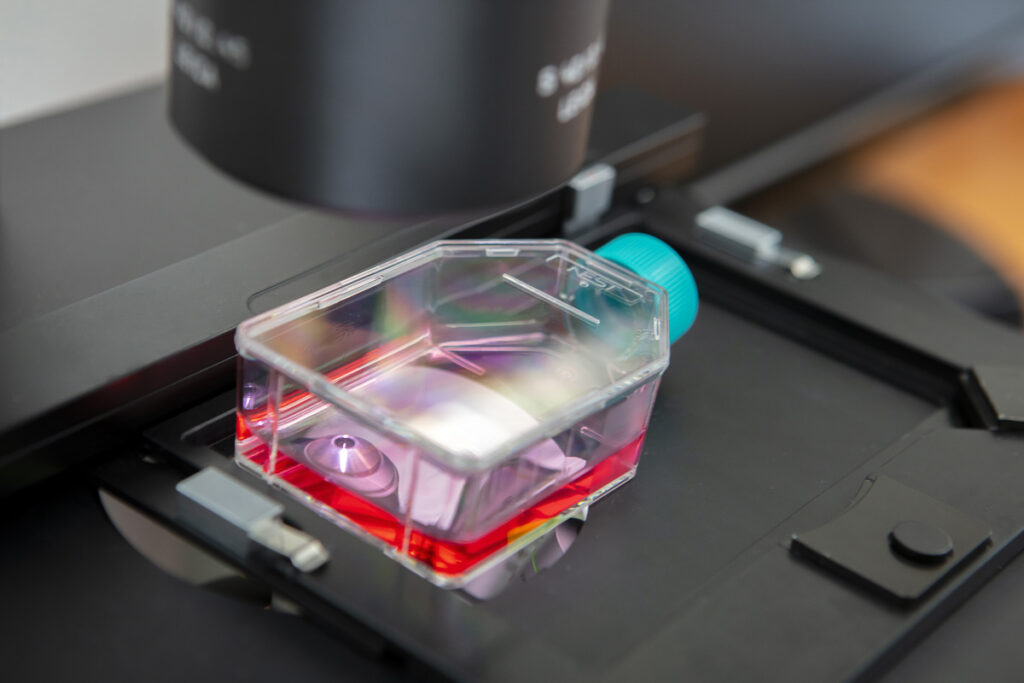

CELL BIOLOGY
Our cell biology platform enables us to handle different eukaryotic cell lines and expose them to microorganisms. Manipulating these cells allows us to offer cytotoxicity and cell viability evaluation services induced by compounds, bacteria, fungal microorganisms, or phages. Additionally, these models enable early-stage study of infection parameters. Eukaryotic cell lines are also useful for studying or producing intracellular bacteria.
In summary, the cell biology platform includes:
- 32 m² laboratory complying with Biosafety Level 2 (BSL2)
- Inverted fluorescence microscope
- Cell banks stored in liquid nitrogen – on a remote platform
- Tecan Spark 10M automated cell counting reader
Our Quality Requirements
These technical platforms are equipped with high-tech equipment validated by the laboratory’s teams, allowing us to provide high-quality services. Smaltis regularly acquires devices to meet the needs of new projects and stay at the forefront of technology.
Smaltis collaborates with qualified suppliers of materials, reagents, and equipment, with whom a relationship of trust has been established, some since the company’s inception. We are committed to using quality and robust equipment, essential for the smooth running of manipulations and the reliability of the results obtained. We carefully select the best to give you the best!
Our laboratory doors are always open; feel free to visit our premises and platforms!
An extensive network of partners and working groups
To provide turnkey solutions, Smaltis has surrounded itself with numerous partners. Smaltis is also a member of various networks to stay in touch with numerous stakeholders in the field of microbiology applied to health and to clearly understand the sector’s needs.
Joining POLEPHARMA means becoming part of a territory of shared needs, expertise, and solutions where everyone contributes to the excellence of all. Joining POLEPHARMA means participating in the economic and industrial network of stakeholders in the leading French pharmaceutical sector: Research laboratories, Production sites, Contract manufacturers, Suppliers, Training centers.
Read more
AFSSI, the French Association of Service and Innovation Companies for Life Sciences, was established in 2012 to bring together French companies providing services and technological innovation in the strategic field of Life Sciences. AFSSI aims to gather all sectors of biotechnology, chemistry, environment, cosmetology, agri-food, bioinformatics, including diagnostics and clinical trials.
Read more
Lyonbiopôle Auvergne-Rhône-Alpes, labeled as a competitiveness cluster at its creation, is the catalyst for the health innovation ecosystem in Auvergne-Rhône-Alpes, ensuring its connection, development, and promotion.
Read more
MabDesign is the French association for the industrial sector of biomedicines. Created in 2014 in response to the "Strategic Future Industrial Sector" call for projects, MabDesign aims to structure the industrial sector of biomedicines in France from R&D phases to market release, including bioproduction, and to generate the creation of innovative start-ups from French academic research.
Read more
PMT, a competitiveness cluster labeled by the French government in 2005, ws established as an Association uner the Law of 1901. Its mission is to catalyze innovation and accelerate the business of industrial companies in Bourgogne-Franche-Comté. PMT supports more than 240 industrial members specialized in specialty subcontracting or developing high-value-added products, offering daily and individualized support.
Read more
A group based on complementary expertise with multidisciplinary know-how. A range of integrated solutions in biotechnology. A complete platform, from DNA to recombinant antibodies. Customized services and products in Biotechnology: Molecular Biology – Immunology – Protein and Cell Engineering. An efficient synergy within a team of experts.
Read more
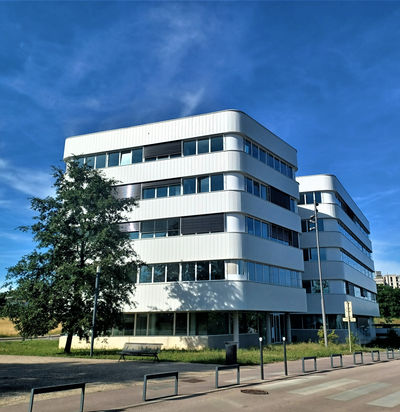

Our location
Smaltis established itself right from its inception in the heart of the TEMIS Santé zone in Besançon, built around the Jean-Minjoz University Hospital Center, the French Blood Establishment (Etablissement Français du Sang), and the National Institute of Health and Medical Research (INSERM). This business park was specifically designed to provide exceptional conditions to launch new ventures and aims to become a national hub for biomedical engineering. It is only logical that Smaltis positioned itself at the center of this dynamic network, aiming to be a genuine participant in ambitious undertaking, while being surrounded by partners focused on research and innovation in health, such as RD-Biotech, Skinexigence, Macopharma, Lymphobank, ISIFC, and Biotika Biomedical.
Get in touch




















